View in English?
View in English?
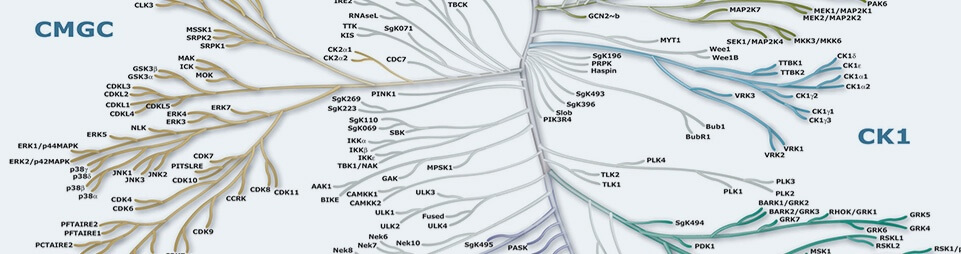
プロテインキナーゼは、細胞機能の重要な調節因子であり、最大かつ最も機能的で多様な遺伝子ファミリーの1つを構成します。このプロテインキナーゼは、基質タンパク質にリン酸化基を付加することで、活性化や局在を調節し、多くのタンパク質の全体的な機能を方向づけを行い、ほぼすべての細胞内プロセスの活性を取り仕切ります。キナーゼは、特にシグナル伝達や細胞周期のような複雑な機能の調整において、卓越した機能を発揮します。酵母から無脊椎動物、ヒトまで、キノームに、50ほどの異なるキナーゼファミリーが保存されていることからも、キナーゼが多様な細胞機能に不可欠であることが分かります。ヒトタンパク質キナーゼ518種のうち478種は、触媒ドメインの配列が関連した単一のスーパーファミリーに属しています。これらのキナーゼは、配列類似性の高さや生化学的機能により、グループ、ファミリー、サブファミリーにクラスター化することが可能です。これらの触媒ドメイン間の配列類似性を示すのが、キナーゼ樹状図 (上) です。2つのキナーゼの間の枝分かれの距離は、この配列の間の相違の程度に比例します。7つの主要なグループがラベルされ、それぞれ色分けされています。例えば、チロシンキナーゼのメンバーは、タンパク質のチロシン残基上をリン酸化する、1つの独立したグループを形成します。一方、その他すべてのグループの酵素は主にセリンやスレオニン残基をリン酸化します。この樹状図上で示される関係は、場合によっては、ここに示されている100以上の性質不明の多くのキナーゼのタンパク質の基質や生物学的機能の予測に使用することもできます。さらに40の「非定型的 (atypical)」キナーゼは、典型的なキナーゼとの配列類似性はありませんが、酵素活性を持つことが知られている、またはそれが推測されており、一部は典型的なキナーゼと類似した折り畳み構造を持つことが推測されています。
Gerard Manning:Sugen, Inc. (米国カリフォルニア州、南サンフランシスコ)