View in English?
View in English?
| Product Includes | Volume (with Count) | ||||
|---|---|---|---|---|---|
| PTMScan® HS Mono-Methyl Arginine (mme-RG) Magnetic Immunoaffinity Beads | 1 x 200 µl | ||||
| PTMScan® IAP Buffer (10X) 9993 | 10 x 600 µl |
Product Information
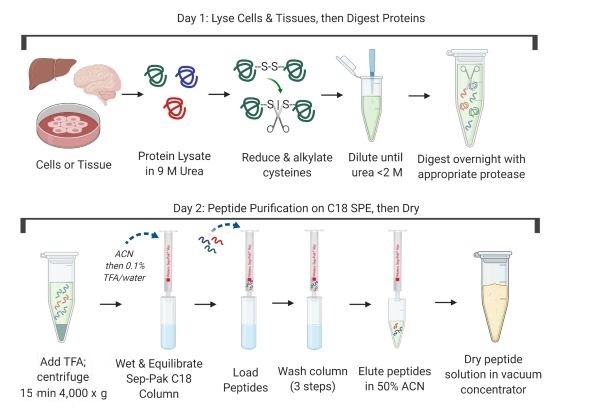
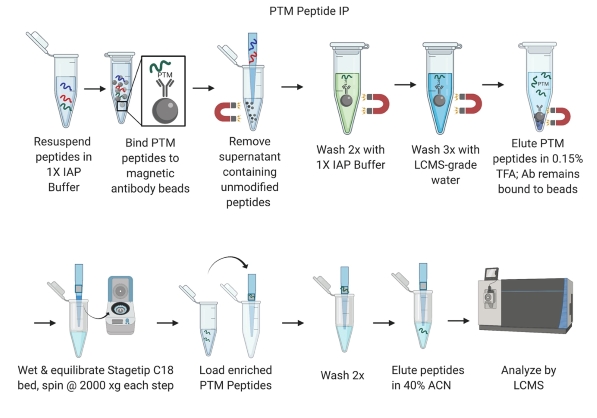
PTMScan enrichment kits are compatible with many protein extraction, digestion, and purification protocols. Compatible workflows include in-solution digestion, those that use centrifugal reactors (FASP1, S-Trap™ cartridges2, or iST3) or magnetic bead precipitation (SP34). Regardless of the particular method selected, ensure that peptides are completely dry and free of lysis buffer components, lipids, and excess salts prior to using the immunoaffinity purification kit. Below is a general protocol that uses in-solution digestion followed by solid phase extraction on Sep-Paks.
References:
NOTE: Prepare solutions for cell lysis (Section I), C18 column purification (Section II), and IAP enrichment (Section III) with reverse osmosis deionized (RODI) or equivalent grade water. Prepare solutions using HPLC grade water (Burdick and Jackson water) for the peptide concentration steps (Section IV).
NOTE: Prepare solutions with RODI or equivalent grade water.
NOTE: The Urea Lysis Buffer should be prepared fresh prior to each experiment. Do not include protease inhibitors.
NOTE: Dissolving urea is an endothermic reaction. Urea Lysis Buffer preparation can be facilitated by placing a stir bar in the beaker and by using a warm (not hot) water bath on a stir plate. 9 M urea is used so that upon lysis, the final concentration is approximately 8 M. The urea lysis buffer should be used at room temperature. Placing the urea lysis buffer on ice will cause the urea to precipitate out of solution.
NOTE: DO NOT place Urea Lysis Buffer or culture dishes on ice during harvesting. Harvest cells using Urea Lysis Buffer at room temperature. During lysis, the buffer becomes viscous due to DNA released from the cells.
NOTE: If desired, the PTMScan® protocol may be interrupted at this stage. The lysed cells or tissues can be frozen and stored at -80°C for several weeks.
NOTE: Centrifugation is performed at room temperature to prevent urea from precipitating out of solution.
NOTE: Lysate sonication fragments DNA and reduces sample viscosity. Ensure that the sonicator tip is submerged in the lysate. If the sonicator tip is not submerged properly, it may induce foaming and degradation of your sample.
NOTE: WaLP is used for SUMOylated peptide analysis in combination with the Ubiquitin/SUMO Remnant Kit (Cell Signaling Technology, #59322). All other PTMScan HS samples should be prepared with trypsin as the protease.
NOTE: Alternative proteases such as GluC, chymotrypsin, and others can be used in addition to the protease digests outlined in the reference table to expand the coverage of modified peptides from each Motif Antibody. When considering the use of additional protease digests it should be compatible with the respective Motif Antibody by not cleaving residues within the designated sequence motif. Alternate protease digests that generate larger proteolytic peptides may not be ideal if the resulting peptides do not ionize well in the mass spectrometer.
NOTE: Use WaLP only for generating KGG remnants in SUMOylated protein experiments.
NOTE: Purification of peptides is performed at room temperature on C18 reversed-phase Sep-Pak columns from Waters (#WAT054955).
NOTE: C18 purification uses reversed-phase (hydrophobic) solid-phase extraction. Peptides and lipids bind to the chromatographic material. Large molecules such as DNA, RNA, and most protein, as well as hydrophilic molecules such as many small metabolites are separated from peptides using this technique. Peptides are eluted from the column with 50% acetonitrile (ACN) and separated from lipids and proteins, which elute at approximately 60% ACN and above.
NOTE: About 2.5 mg of protease-digested peptides can be purified from one C18 column. Purify peptides immediately after proteolytic digestion.
NOTE: Prepare solutions with HPLC grade water. Pierce Trifluoroacetic Acid (TFA), Sequencing grade (ThermoFisher Scientific, 28903) and Pierce Acetonitrile (ACN), LC-MS Grade (Thermo Scientific, 51101) when preparing solutions. All percentage specifications for solutions are vol/vol.
NOTE: Organic solvents are volatile. Tubes containing small volumes of these solutions should be prepared immediately before use and should be kept capped as much as possible because the organic components evaporate quickly.
NOTE: Before loading the peptides from the digested sample on the column, they must be acidified with TFA for efficient peptide binding. The acidification step helps remove fatty acids from the digested peptide mixture.
NOTE: Application of all solutions can be performed with a vacuum manifold or by gravity flow. If using vacuum, keep flow rates below approximately 0.33mL/min for most steps. Sample loading should be done by gravity flow to maximize recovery.
NOTE: Peptide solutions may be frozen at -80 °C for 1 hr or longer before placing in the Speed-Vac; this will prevent full tubes from spilling when placed at an angle to dry.
NOTE: A standard lyophilization apparatus is also acceptable in place of a vacuum concentrator.
NOTE: Dry, digested peptides are stable at -80°C for several months (seal the closed tube with parafilm for storage). The PTMScan® procedure can be interrupted before or after drying. Once the dry peptide is dissolved in 1X IAP Buffer (see next step), continue to the end of the procedure.
NOTE: Tubes can be shaken gently at room temperature using a vortexer or thermomixer for 5 min or placed in a sonicator bath for 2 min to ensure complete solubilization, if necessary.
NOTE: After dissolving the peptide, check the pH of the peptide solution by spotting a small volume on pH indicator paper. The pH should be close to neutral (no lower than 7.0). If necessary, add 2 µL of 1M Tris base at a time until the pH is at 7.0.
NOTE: There may be a small, insoluble pellet. Transfer supernatant to a clean tube and discard the pellet.
NOTE: Ensure the beads remain in suspension while rotating and that bubbles do not collect at the bottom of the tube as this will prevent proper bead and sample mixing.
NOTE: Keep the 1X IAP Buffer and LCMS Water on ice for the subsequent steps.
NOTE: In this step, the post-translationally modified peptides of interest will be in the eluent.
NOTE: We recognize there are many routine methods for concentrating peptides using commercial products such as C18 tips (see below) that have been optimized for peptide desalting/concentration. Regardless of the particular method, we recommend that the method of choice be optimized for recovery and be amenable for peptide loading capacities of at least 10 µg.
C18 tips: Pierce C18 Spin Tips (ThermoFisher Scientific 84850)
NOTE: Prepare solutions with Burdick and Jackson water or other LCMS grade water. Organic solvents (trifluoroacetic acid, acetonitrile) should be of the highest grade.
Recommended: Pierce Trifluoroacetic Acid (TFA), Sequencing grade (ThermoFisher Scientific, 28903) and Pierce Acetonitrile (ACN), LCMS Grade (ThermoFisher Scientific, 51101).
Prepare all solutions in containers that have never been exposed to soap, as detergents will interfere with LCMS analysis.
NOTE: Organic solvents are volatile. Tubes containing small volumes of these solutions should be prepared immediately before use and should be kept capped as much as possible, to prevent evaporation of organic components.
NOTE: All centrifugation steps in this section should be carried out at room temperature. Spin at 2,000 x g or a speed that passes all the solution through the tip in approximately 3 min.
Protocol Id: 2704
Arginine methylation is a prevalent PTM found on both nuclear and cytoplasmic proteins. Arginine methylated proteins are involved in many different cellular processes, including transcriptional regulation, signal transduction, RNA metabolism, and DNA damage repair (1-3). Arginine methylation is carried out by the arginine N-methyltransferase (PRMT) family of enzymes that catalyze the transfer of a methyl group from S-adenosylmethionine (AdoMet) to a guanidine nitrogen of arginine (4). There are three different types of arginine methylation: asymmetric dimethylarginine (aDMA, omega-NG,NG-dimethylarginine), where two methyl groups are placed on one of the terminal nitrogen atoms of the guanidine group of arginine; symmetric dimethylarginine (sDMA, omega-NG,NG-dimethylarginine), where one methyl group is placed on each of the two terminal guanidine nitrogens of arginine; and monomethylarginine (MMA, omega-NG-methylarginine), where a single methyl group is placed on one of the terminal nitrogen atoms of arginine. Each of these modifications has potentially different functional consequences. Though all PRMT proteins catalyze the formation of MMA, Type I PRMTs (PRMT1, 3, 4, 6, and 8) add an additional methyl group to produce aDMA, while Type II PRMTs (PRMT5 and 7) produce sDMA. Methylated arginine residues often reside in glycine-arginine rich (GAR) protein domains, such as RGG, RG, and RXR repeats (5). However, PRMT4/CARM1 and PRMT5 methylate arginine residues within proline-glycine-methionine rich (PGM) motifs (6).
Except as otherwise expressly agreed in a writing signed by a legally authorized representative of CST, the following terms apply to Products provided by CST, its affiliates or its distributors. Any Customer's terms and conditions that are in addition to, or different from, those contained herein, unless separately accepted in writing by a legally authorized representative of CST, are rejected and are of no force or effect.
Products are labeled with For Research Use Only or a similar labeling statement and have not been approved, cleared, or licensed by the FDA or other regulatory foreign or domestic entity, for any purpose. Customer shall not use any Product for any diagnostic or therapeutic purpose, or otherwise in any manner that conflicts with its labeling statement. Products sold or licensed by CST are provided for Customer as the end-user and solely for research and development uses. Any use of Product for diagnostic, prophylactic or therapeutic purposes, or any purchase of Product for resale (alone or as a component) or other commercial purpose, requires a separate license from CST. Customer shall (a) not sell, license, loan, donate or otherwise transfer or make available any Product to any third party, whether alone or in combination with other materials, or use the Products to manufacture any commercial products, (b) not copy, modify, reverse engineer, decompile, disassemble or otherwise attempt to discover the underlying structure or technology of the Products, or use the Products for the purpose of developing any products or services that would compete with CST products or services, (c) not alter or remove from the Products any trademarks, trade names, logos, patent or copyright notices or markings, (d) use the Products solely in accordance with CST Product Terms of Sale and any applicable documentation, and (e) comply with any license, terms of service or similar agreement with respect to any third party products or services used by Customer in connection with the Products.